Staphylococcus epidermidis: A Potential New Player in the Physiopathology of Acne?
This article was published in the Vol 23, No. 4, 2019 Issue of Karger. We would like to thank Dr Jean-Paul Claudel, Dr Nicole Auffret, Dr Marie-Therese Lecciac, Dr Florence Poli, Dr Stephane Corvec, Dr Brigitte Dreno and Karger for sharing it with the international community.
Keywords Acne vulgaris · Acne · Cutibacterium acnes · Dysbiosis · Microbiota · Pilosebaceous unit · Sebum · Staphylococcus epidermidisAbstract
Background: Cutibacterium acnes has been identified as one of the main triggers of acne. However, increasing knowledge of the human skin microbiome raises questions about the role of other skin commensals, such as Staphylococcus epidermidis, in the physiopathology of this skin disease. Summary: This review provides an overview of current knowledge of the potential role of S. epidermidis in the physiopathology of acne. Recent research indicates that acne might be the result of an unbalanced equilibrium between C. acnes and S. epidermidis, according to dedicated interactions. Current treatments act on C. acnes only. Other treatment options may be considered, such as probiotics derived from S. epidermidis to restore the naturally balanced microbiota or through targeting the regulation of the host’s AMP mediators. Key Messages: Research seems to confirm the beneficial role of S. epidermidis in acne by limiting C. acnes over-colonization and inflammation.
Introduction
With the availability of bioinformatic tools leading to new phylogenetic approaches, important advances have been made over the last years either to investigate the diversity and the topology of skin microbiota or to identify the different commensal microorganisms present on the skin, to evaluate the relative abundance of each population, and to understand their beneficial role or contribution to dermatological conditions such as acne [4-6]. Metagenomic analysis and sequencing of the 16S ribosomal RNA gene are the predominant bacteriological sampling methods employed for analysing the bacterial composition of microbial communities, and choosing the most effective study design is crucial for obtaining meaningful analysis results [7]. Thus, physiological characteristics of different skin sites sampled have been associated with different levels of bacterial diversity including Actinobacteria such as Propionibacteria, very recently renamed Cutibacteria (comprising Cutibacterium acnes and also Cutibacterium granulosum and Cutibacterium avidum), Proteobacteria, Bacteroides and Firmicutes; Staphylococcus epidermidis belongs to the latter group [5, 6, 8, 9]. Of these, C. acnes and S. epidermidis are two of the main commensal skin bacteria [3, 10-12]. Until very recently, research in acne has mainly focused on the role of C. acnes while the role of S. epidermidis in acne has been discussed for several years but still remains to be elucidated [10, 13].
The aim of the present article is to review published data concerning the role of S. epidermidis in the physiopathology of acne and to present future treatment perspectives based on available data and evidence.
Methodology
A review of the literature published between the beginning of 2000 and 2018 and available from PubMed was carried out using the following key words: acne, acne vulgaris + microbiota, skin + microbiota, acne + microbiota + Propionibacterium acnes + Staphylococcus epidermidis, bacterial resistance, skin commensal + Propionibacterium acnes + Staphylococcus epidermidis.
Results The Interplay of Host Skin and the Cutaneous Microbiota
The interplay of host skin and the cutaneous microbiota in the skin’s immune system is able to differentiate between harmless commensal microorganisms including Corynebacterium sp., Staphylococcus sp. excluding S. aureus, Cutibacterium sp., Malassezia furfur, and transient, harmful pathogenic microorganisms such as S. aureus, Streptococcus pyogenes, and Enterobacteriaceae. Even though it seems as if this recognition involves the host’s immune response, this mechanism is not fully understood. Toll-like receptors (TLRs) have been reported to be desensitised through prolonged exposure to commensal microorganisms, either through a decreased expression of TLR on the cell surface or by activation of the TLR pathway inhibitors interleukin-1 receptor-associated kinase 3 (IRAK3) and the suppressor of cytokine signalling 1 (SOCS1) [14, 15]. Moreover, specificity may be achieved by a combined recognition of pathogen-associated molecular patterns through pattern recognition receptors [6].
Thus, both such recognised commensals C. acnes and S. epidermidis seem to interact with the host, helping to protect the healthy skin from colonisation by pathogens [9, 16]. Figure 1 provides visual support.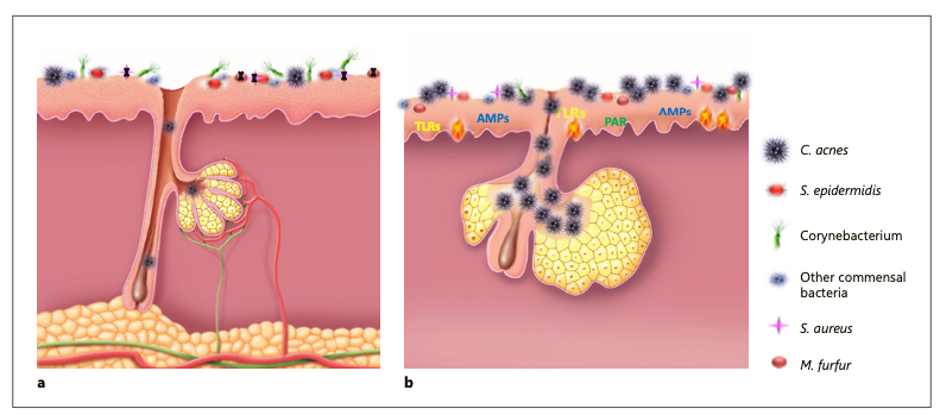
Fig. 1. Healthy skin and skin with acne lesion in formation. a Healthy skin. In healthy skin S. epidermidis controls the proliferation of C. acnes. b Formation of a microcomedone after over-colonisation of the skin by C. acnes leading to dysbiosis. Over-colonisation of C. acnes during puberty leads to dysbiosis and acne. AMP, antimicrobial peptides; TLR, Toll-like receptors; PAR, protease-activated receptors.
However, the composition of the cutaneous microbiota evolves constantly and changes over time. As an example, a disturbed balance or dysbiosis of previously healthy skin caused by exogenous factors such as injury, stress, or pollution, or endogenous factors (hormonal changes, changes of pH) may induce inflammatory skin disorders such as acne, atopic dermatitis, rosacea, and psoriasis [13, 17-21]. Specifically, during puberty, the increase and qualitative modification of sebum due to hormonal modifications may lead to an unbalanced skin microbiota via an increased colonisation of the pilosebaceous unit and of the skin surface by C. acnes[6]. This over-representation may be due to a selective advantage linked to the nutritive substance availability [22].
The Role of C. acnes
In healthy skin, C. acnes plays a beneficial role in the cutaneous microbiota of the pilosebaceous unit. It limits the growth of S. aureus, such as community-acquired methicillin-resistant S. aureus, as well as that of S. pyogenes in maintaining an acidic pH of the pilosebaceous follicle by hydrolysing sebum triglycerides and by secreting propionic acid [6, 23, 24].
However, during puberty, over-colonisation of the pilosebaceous unit by C. acnes may lead to a loss of diversification and dysbiosis, potentially causing acne [6, 25-27]. A recent clinical study using the single-locus sequence typing method investigated C. acnes subgroups on the face and back in patients with severe acne and in healthy subjects [28]. In almost 75% of the acne patients, C. acnes phylotypes were identical on the face and back, whereas this was only the case in about 45% of the healthy subjects. In the healthy group, phylotypes IA1 (39%) and II (43%) were the main phylotypes, whereas in the acne group IA1 (84%), especially on the back (96%), was the main phylotype. This may confirm the hypothesis that acne severity may be associated with a loss of diversity of C. acnes phylotypes, following a selection of phylotype IA1/clonal complex (CC) 18 present in all acne patients [28-30].
Therefore,different inflammatory profiles, depending on the phylotype (i.e., phylotype IA1, which has been mainly observed on the face and back of acne patients and clusterof C. acnes activating the innate immunity via the expression of protease-activated receptors (PARs), tumour necrosis factor-a, and the production of interferon-y and interleukins [IL-8], have been observed [28, 31 -37]. Moreover, C. acnes activates the release of lipases, matrix metalloproteinases, and hyaluronidases, leading to hyperkeratinisation of the pilosebaceous unit and finally to comedones, papules, and pustules [31-34, 38].Figure 1 shows the difference between healthy microbiome and microbiome suffering from dysbiosis.
The Role of S. epidermidis
S. epidermidis is the most frequently isolated commensal species from human epithelia [25, 39]. It colonises predominantly axillae, the head, and nares, and has generally a non-pathogenic relationship with its host [11, 39].
S. epidermidis belongs to the group of coagulase-negative staphylococci, which differs from coagulase-positive staphylococci such as S. aureus by lacking the enzyme coagulase. According to multi-locus sequence typing, the species showed a high degree of diversity with more than 400 identified sequence types (STs) compared to 155 for C. acnes[40, 41]. Most clinical isolates belong to CC2, which comprises the most frequently isolated ST2. Possibly, the successful spread of ST2 may be due to the fact that all ST2 isolates contain IS256 insertion sequences and ica genes, two factors found to be correlated with S. epidermidis invasiveness [42, 43]. Nevertheless, controversy still exists about the fact that icaA is a useful marker for S. epidermidis biofilm [44].
In the past, S. epidermidis has been considered an innocuous commensal microorganism on the human skin. Nowadays it is seen as an important opportunistic pathogen. It is the most frequent cause of nosocomial infections, at a rate about as high as that due to its more virulent cousin S. aureus. In particular, S. epidermidis represents the most common source of infection on indwelling medical devices [45].
Treatment is complicated by specific antibiotic-resistant genes and the formation of biofilms, multicellular agglomerations that have intrinsic resistance and tolerance to antibiotics and mechanisms of host defence [46]. Furthermore, investigations have identified specific molecular determinants allowing S. epidermidis immune invasion and its ability to cause chronic diseases [47]. These causes are believed to have original functions in the non-infectious lifestyle of this microorganism, emphasising the accidental nature of S. epidermidis infections allowing differentiating between portage, contamination, and infection. A better understanding of the physiology of S. epidermidis is important to evaluate therapeutic strategies against S. epidermidis infections [39].
Certain strains of S. epidermidis may modulate the host innate immune response, especially of TLR 2, and thus allow the host to fight against pathogens [6, 28]. Phenol-soluble modulins produced by S. epidermidis were shown to selectively inhibit skin pathogens, such as S. aureus and Group A Streptococcus, and, together with host antimicrobial peptides (AMPs), enhanced their elimination [48-50]. Moreover, S. epidermidis activates the AMP expression of keratinocytes through a TLR2-dependent mechanism [51]. This confirms that this skin commensal interacts closely with the host innate immune mechanism [48, 49]. This was also shown through the microdissection of epidermal compartments of the dermis and of the adipose tissue followed by 16S ribosomal RNA sequencing demonstrating that colonisation with S. epidermidis tunes T-cell homing and function in an IL-1-dependent manner [52, 53].
Other very recent research work reported that S. epidermidis may also protect against skin neoplasia through the production of 6-N-hydroxyaminopurine, a molecule that inhibits DNA polymerase activity [54]. Moreover, butyric acid, released by S. epidermidis, allows adipose-derived stem cells to differentiate into adipocytes and lipid accumulation in the cytoplasm, resulting in an increased dermal layer [55].
The Interplay between S. epidermidis and C. acnes in Acne
S. epidermidis and C. acnes use glycerol as a shared carbon source to produce different short-chain fatty acids (SCFAs) used as antimicrobial agents to compete against each other. Theyare both present in acne lesions. This concomitant presence has raised questions about their respective role in the pathogenesis of acne [56, 57]. While there is no evidence that S. epidermidis plays an active role in the onset of acne, C. acnes is currently associated with acne [9].
Some S. epidermidis isolates have been shown through an antagonism assay to possess antimicrobial activity against C. acnes [27, 56]. Among the S. epidermidis strains with an elevated antimicrobial activity, differences in inhibition zone diameter and appearance were observed, indicating that the antimicrobial substances were of a different nature. Most of the S. epidermidis strains displayed small zones of inhibition (2–4 mm) against C. acnes and some strains produced opaque zones of inhibition. One strain, FS1, produced very large inhibitory zones (>10 mm), but was not active against all C. acnes strains tested. Another strain, 14.1.R1, inhibited all C. acnes strains, but produced only small zones of inhibition (2–5 mm). No difference in prevalence and intensity was noted in the antimicrobial activity of S. epidermidis strains isolated from normal and acne-affected skin, respectively. Likewise, the origin of C. acnes strains did not determine their susceptibility to the antimicrobial activity of S. epidermidis, since strains isolated from acne lesions and healthy skin, respectively, were not significantly different in their susceptibility patterns towards S. epidermidis. Conversely, the study showed a higher frequency of antimicrobial activity among certain C. acnes strains, such as I-2 phylogroup, against S. epidermidis. The authors hypothesised a likely possible presence and secretion of a bacteriocin or bacteriocin-like substance specific to C. acnes phylogroup I-2 [13]. C. acnes strains and their antimicrobial activities against S. epidermidis have also been investigated. However, regarding the skin status origin (healthy, moderate, or severe acne), the antimicrobial activity ranged from 29 to 39%. Consequently, type IA1 CC18 C. acnes strains largely involved in acne did not present higher activity compared to healthy strains [13].
Different antagonism investigations demonstrated that, in vivo, S. epidermidis controls the proliferation of C. acnes via the release of succinic acid, a fatty acid fermentation product, which inhibits surface TLRs of keratinocytes and tumour necrosis factor and suppresses C. acnes-induced IL-6 [27, 56, 58].
With both C. acnes and S. epidermidis being present on the skin, the inhibition of C. acnes- induced inflammation by S. epidermidis may potentially be dependent on staphylococcal LTA-induced miR-143 on keratinocytes, known for limiting inflammation. Research suggested that the mechanism for LTA-miR-143-mediated suppression of TLR2 signalling is accomplished by miR-143 targeting the 3'UTR of TLR2. It thereby decreases the TLR2 protein production, which plays a major role in the inhibition of C. acnes-induced cutaneous inflammation [58-60]. Thus, it helps to regulate skin homeostasis and to suppress the pathogenic inflammation that is induces by C. acnes [58-61]
Accordingly, an unbalanced equilibrium between C. acnes and S. epidermidis in pilosebaceous units of acne patients in favour of phylotype IA1 CC18 C. acnes strains (75–80 of cases) may not allow S. epidermidis to fully play its role as a regulator of the natural skin homeostasis in limiting the growth of C. acnes. Figure 2 shows visually the interplay between C. acnes and S. epidermidis.
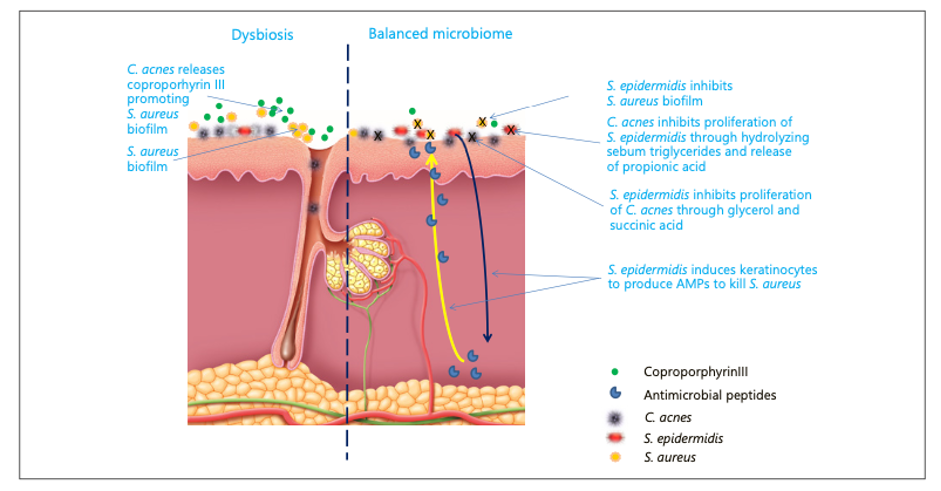
Fig. 2 Bacterial interactions on the skin. AMP, antimicrobial peptides.
S. epidermidis and its Implication in the Choice of Current and Future Acne Treatments
Current topical acne treatments principally include different retinoids, such as adapalene and tretinoin, which reduce inflammation by modulating the innate immunity activated by C. acnes [62-65]. Apart from benzoyl peroxide, the use of topical antibacterials such as erythromycin and clindamycin in monotherapy resulted in the development of antibiotic-resistant strains of not only C. acnes but also of S. epidermidis in 4–6 weeks [30, 57, 66]. Systemic antibiotics have been reported to cause only little resistance but can lead to collateral damage in gut microbiota [30, 33, 67]. Oral isotretinoin normalises the response of the innate immune system to C. acnes by inhibiting its proliferation [68].
Thus, eliminating only C. acnes may favour the proliferation of S. aureus, triggering inflammatory acne flare-up and proliferation of S. epidermidis, leading to another unbalanced skin homeostasis and a risk of nosocomial infections.
AMPs are effector molecules of the innate immune system of the skin. They are amphipathic and disrupt the lipid membrane of the bacterium, leading to cell lysis and death by interacting preferentially with negatively charged bacterial membranes rather than with neutrally charged mammalian cell membranes. The importance of the AMPs’ contribution to host immunity is undeniable as alterations in AMP expression have been associated with various pathologic processes. Nevertheless, data regarding the role of AMPs in acne vulgaris are limited. A recently published study reports that the AMPs hBD-1 and cathelicidins play an important role in the pathogenesis of acne [69]. AMPs have shown activity against a broad spectrum of Gram-positive and Gram-negative bacteria, as well as some fungi, parasites, and enveloped viruses [70]. They are produces asa response by keratinocytes and sebocytes. However, they also promote additional inflatammory responses. The production of AMPs comprises B-defensins, RNase 7, the S100-protein psoriasin, and cathelicidins, and is mediated by the MyD88 pathway and IL-1 signalling. In addition, it has been demonstrated that more cells express TLR-2 as acne severity increases [33]. This may be one explanation why agents that target TLR-2, such as topical retinoids, have a greater efficacy in patients with more sever acne [71]. Cytokines are also produced as a result of the interaction between C. acnes and TLR-2, defensins, and matrix metalloproteinase via PAR-2R activation [72].
Another treatment approach could be the regular oral or topical supplementation of the skin microbiota with a beneficial microorganism (probiotics) for patients with acne by re-equilibrating the cutaneous microbiota [73, 74]. In 2010, Arck et al. [75] suggested that not only a brain-gut or brain-skin axis, but also a gut-skin axis exists. The authors showed that oral supplementation of a Lactobacillus strain in mice dampens stress-induced neurogenic skin inflammation and hair growth inhibition. Their concept suggested that modulation of the microbiota by deployment of probiotics reduces stress-induced neurogenic skin inflammation. These observations raise hope that the right kind of bacteria may have beneficial effects on the skin homoeostasis, skin inflammation, and peripheral tissue responses to perceived stress. Other studies conducted in humans confirmed this hypothesis [76-78]. Thus, probiotics may be efficient in acne and other inflammatory skin diseases such as atopic dermatitis and potentially psoriasis [79-83].
In this perspective, studies indicate that S. epidermidis may control dysbiosis caused by C. acnes, thus reducing acne severity [27, 56]. However, it still remains unclear which SCFA in the products of S. epidermidis glycerol fermentation primarily contributes to the anti-C. acnes effect. It is also undetermined whether SCFAs act together with other antimicrobial molecules in fermentation products to display their anti-C. acnes activities. The anti-C. acnes activity of the fermented media persisted after boiling the said media, thereby suggesting that the antimicrobial proteins/peptides may not be the major contributors to the anti-C. acnes activity of fermented media [56].
Pro-drugs such as the SCFAs pivaloylomethyl butyrate (AN-9) have been developed to achieve pharmacologic concentrations of such SCFAs in vivo[84]. Moreover, live S. epidermidis may potentially be used as an active component in acne probiotics [27, 56].
Discussion and Conclusion
Acne is a chronic and multifactorial inflammatory disease of the skin and the pilosebaceous unit. Dysbiosis in acne patients is associated with a decreased number of S. epidermidis and an over-colonisation by selected C. acnes phylotypes in the sebaceous unit, leading to different levels of activation of the innate immunity, thus resulting in different severity levels of inflammatory acne. Recent research seems to confirm the beneficial role of S. epidermidis in the physiopathology of acne through limiting C. acnes-induced colonisation of the skin and of inflammation [9, 33]. However, disequilibrium in favour of S. epidermidis may also result in other health consequences such as nosocomial infections. Therefore, a balanced skin homeostasis should be the final aim in any acne treatment.
In this perspective, other treatment options may be considered, such as probiotics derived from S. epidermidis, allowing for a restoration of the naturally balanced microbiota, and through the targeting of the regulation of the host’s AMP mediators, without increasing the local S. epidermidis population.
Key Message S. epidermidis limits C. acnes-induced colonisation of the skin, thus potentially playing a role in the physiopathology of acne.
Acknowledgements The authors acknowledge the writing support of Patrick Göritz, SMWS-Scientific, and Medical Writing Services.
Statement of Ethics The authors have no ethical conflicts to disclose
Disclosure Statement The authors have no financial interest to disclose.
Funding Sources This Expert Group was organised by Galderma International, France.
References
1. Kong HH, Segre JA. Skin microbiome: look- ing back to move forward. J Invest Dermatol. 2012 Mar;132(3 Pt 2):933–9.
2. Cogen AL, Nizet V, Gallo RL. Skin microbio- ta: a source of disease or defence? Br J Derma- tol. 2008 Mar;158(3):442–55.
3. Oh J, Byrd AL, Park M, Kong HH, Segre JA; NISC Comparative Sequencing Program. Temporal Stability of the Human Skin Microbiome. Cell. 2016 May; 165(4): 854–66.
4. Peterson J, Garges S, Giovanni M, McInnes P, Wang L, Schloss JA, et al.; NIH HMP Work- ing Group. The NIH Human Microbiome Project. Genome Res. 2009 Dec;19(12):2317– 23.
5. Grice EA, Kong HH, Conlan S, Deming CB, Davis J, Young AC, et al.; NISC Comparative Sequencing Program. Topographical and temporal diversity of the human skin micro- biome. Science. 2009 May;324(5931):1190– 2.
6. Grice EA, Segre JA. The skin microbiome. Nat Rev Microbiol. 2011 Apr;9(4):244–53.
7. Meisel JS, Hannigan GD, Tyldsley AS, San- Miguel AJ, Hodkinson BP, Zheng Q, et al. Skin Microbiome Surveys Are Strongly Influ- enced by Experimental Design. J Invest Der- matol. 2016 May;136(5):947–56.
8. Scholz CF, Kilian M. The natural history of cutaneous propionibacteria, and reclassifica- tion of selected species within the genus Pro- pionibacterium to the proposed novel genera Acidipropionibacterium gen. nov., Cutibac- terium gen. nov. and Pseudopropionibacte- rium gen. nov. Int J Syst Evol Microbiol. 2016 Nov;66(11):4422–32.
9. Byrd AL, Belkaid Y, Segre JA. The human skin microbiome. Nat Rev Microbiol. 2018 Mar; 16(3):143–55.
10. Coates R, Moran J, Horsburgh MJ. Staphylo- cocci: colonizers and pathogens of human skin. Future Microbiol. 2014;9(1):75–91.
11. Christensen GJ, Bruggemann H. Bacterial skin commensals and their role as host guardians. Benef Microbes. 2014 Jun;5(2):201–15.
12. Fischbach MA, Segre JA. Signaling in Host- Associated Microbial Communities. Cell. 2016 Mar;164(6):1288–300.
13. Christensen GJ, Scholz CF, Enghild J, Rohde H, Kilian M, Thurmer A, et al. Antagonism between Staphylococcus epidermidis and Propionibacterium acnes and its genomic ba- sis. BMC Genomics. 2016 Feb;17(1):152.
14. Fukao T, Koyasu S. PI3K and negative regulation of TLR signaling. Trends Immunol. 2003 Jul;24(7):358–63.
15. Strober W. Epithelial cells pay a Toll for protection. Nat Med. 2004 Sep;10:898–900.
16. Burian M, Bitschar K, Dylus B, Peschel A, Schittek B. The Protective Effect of Microbiota on S. aureus Skin Colonization Depends on the Integrity of the Epithelial Barrier. J Invest Dermatol. 2017 Apr;137(4):976–9.
17. van Rensburg JJ, Lin H, Gao X, Toh E, Fortney KR, Ellinger S, et al. The Human Skin Micro- biome Associates with the Outcome of and Is Influenced by Bacterial Infection. MBio. 2015 Sep;6(5):e01315–15.
18. Scanlan PD, Buckling A, Kong W, Wild Y, Lynch SV, Harrison F. Gut dysbiosis in cystic fibrosis. J Cyst Fibros. 2012 Sep;11(5):454–5.
19. Zeeuwen PL, Kleerebezem M, Timmerman HM, Schalkwijk J. Microbiome and skin dis- eases. Curr Opin Allergy Clin Immunol. 2013 Oct;13(5):514–20.
20. Picardo M, Ottaviani M. Skin microbiome and skin disease: the example of rosacea. J Clin Gastroenterol. 2014 Nov-Dec;48 Suppl 1:S85–6.
21. Prescott SL, Larcombe DL, Logan AC, West C, Burks W, Caraballo L, et al. The skin microbiome: impact of modern environments on skin ecology, barrier integrity, and system- ic immune programming. World Allergy Or- gan J. 2017 Aug;10(1):29.
22. McGinley KJ, Webster GF, Leyden JJ. Region- al variations of cutaneous propionibacteria. Appl Environ Microbiol. 1978 Jan;35(1):62–6.
23. Shu M, Wang Y, Yu J, Kuo S, Coda A, Jiang Y, et al. Fermentation of Propionibacterium acnes, a commensal bacterium in the human skin microbiome, as skin probiotics against methicillin-resistant Staphylococcus aureus. PLoS One. 2013;8(2):e55380.
24. Tax G, Urban E, Palotas Z, Puskas R, Konya Z, Biro T, et al. Propionic Acid Produced by Propionibacterium acnes Strains Contributes to Their Pathogenicity. Acta Derm Venereol. 2016 Jan;96(1):43–9.
25. Bek-Thomsen M, Lomholt HB, Kilian M. Acne is not associated with yet-uncultured bacteria. J Clin Microbiol. 2008 Oct;46(10): 3355–60.
26. Di Domizio J, Pagnoni A, Huber M, Hohl D, Gilliet M. [The skin microbiota: a colossus steps into the spotlight]. Rev Med Suisse. 2016 Mar;12(512):660–4.
27. Wang Y, Kao MS, Yu J, Huang S, Marito S, Gallo RL, et al. A Precision Microbiome Ap- proach Using Sucrose for Selective Augmen- tation of Staphylococcus epidermidis Fer- mentation against Propionibacterium acnes. Int J Mol Sci. 2016 Nov;17(11):E1870.
28. Dagnelie MA, Corvec S, Saint-Jean M, Bourdes V, Nguyen JM, Khammari A, et al. Decrease in Diversity of Propionibacterium acnes Phylotypes in Patients with Severe Acne on the Back. Acta Derm Venereol. 2018 Feb; 98(2):262–7.
29. Aubin GG, Portillo ME, Trampuz A, Corvec S. Propionibacterium acnes, an emerging pathogen: from acne to implant-infections, from phylotype to resistance. Med Mal Infect. 2014 Jun;44(6):241–50.
30. Nakase K, Nakaminami H, Takenaka Y, Hayashi N, Kawashima M, Noguchi N. Relationship between the severity of acne vulgaris and antimicrobial resistance of bacteria isolated from acne lesions in a hospital in Japan. J Med Microbiol. 2014 May;63(Pt 5):721–8.
31. Zouboulis CC, Jourdan E, Picardo M. Acne is an inflammatory disease and alterations of se- bum composition initiate acne lesions. J Eur Acad Dermatol Venereol. 2014 May;28(5): 527–32.
32. Picardo M, Ottaviani M, Camera E, Mastro-francesco A. Sebaceous gland lipids. Dermatoendocrinol. 2009 Mar;1(2):68–71.
33. Dreno B, Gollnick HP, Kang S, Thiboutot D, Bettoli V, Torres V, et al.; Global Alliance to Improve Outcomes in Acne. Understanding innate immunity and inflammation in acne: implications for management. J Eur Acad Dermatol Venereol. 2015 Jun;29 Suppl 4:3– 11.
34. Dreno B. What is new in the pathophysiology of acne, an overview. J Eur Acad Dermatol Venereol. 2017 Sep;31 Suppl 5:8–12.
35. Yu Y, Champer J, Agak GW, Kao S, Modlin RL, Kim J. Different Propionibacterium acnes Phylotypes Induce Distinct Immune Responses and Express Unique Surface and Se- creted Proteomes. J Invest Dermatol. 2016 Nov;136(11):2221–8.
36. Nagy I, Pivarcsi A, Koreck A, Szell M, Urban E, Kemeny L. Distinct strains of Propionibac- terium acnes induce selective human beta-de fensin-2 and interleukin-8 expression in hu- man keratinocytes through toll-like recep- tors. J Invest Dermatol. 2005 May;124(5): 931–8.
37. Jasson F, Nagy I, Knol AC, Zuliani T, Kham- mari A, Dreno B. Different strains of Propi-onibacterium acnes modulate differently the cutaneous innate immunity. Exp Dermatol. 2013 Sep;22(9):587–92.
38. Dessinioti C, Antoniou C, Katsambas A. Ac- neiform eruptions. Clin Dermatol. 2014 Jan- Feb;32(1):24–34.
39. Otto M. Staphylococcus epidermidis—the ‘accidental’ pathogen. Nat Rev Microbiol. 2009 Aug;7(8):555–67.
40. Multi Locus Sequence Typing. Staphylococcus epidermidis. 2018. Available from: http:// sepidermidis.mlst.net/.
41. Meric G, Mageiros L, Pensar J, Laabei M, Ya- hara K, Pascoe B, et al. Disease-associated genotypes of the commensal skin bacterium Staphylococcus epidermidis. Nat Commun. 2018 Nov;9(1):5034.
42. Li M, Wang X, Gao Q, Lu Y. Molecular characterization of Staphylococcus epidermidis strains isolated from a teaching hospital in Shanghai, China. J Med Microbiol. 2009 Apr; 58(Pt 4):456–61.
43. Yao Y, Sturdevant DE, Otto M. Genomewide analysis of gene expression in Staphylococcus epidermidis biofilms: insights into the patho-physiology of S. epidermidis biofilms and the role of phenol-soluble modulins in formation of biofilms. J Infect Dis. 2005 Jan;191(2):289– 98
44. Frank KL, Hanssen AD, Patel R. icaA is not a useful diagnostic marker for prosthetic joint infection. J Clin Microbiol. 2004 Oct;42(10): 4846–9.
45. National Nosocomial Infections Surveillance System. National Nosocomial Infections Sur- veillance (NNIS) System Report, data sum- mary from January 1992 through June 2004, issued October 2004. Am J Infect Control. 2004 Dec;32(8):470–85.
46. Costerton JW, Stewart PS, Greenberg EP. Bacterial biofilms: a common cause of persistent infections. Science. 1999 May;284(5418): 1318–22.
47. Nakamizo S, Egawa G, Honda T, Nakajima S, Belkaid Y, Kabashima K. Commensal bacteria and cutaneous immunity. Semin Immunopathol. 2015 Jan;37(1):73–80.
48. Cogen AL, Yamasaki K, Sanchez KM, Dorschner RA, Lai Y, MacLeod DT, et al. Selective antimicrobial action is provided by phenol-soluble modulins derived from Staphylococcus epidermidis, a normal resident of the skin. J Invest Dermatol. 2010 Jan;130(1): 192–200.
49. Cogen AL, Yamasaki K, Muto J, Sanchez KM, Crotty Alexander L, Tanios J, et al. Staphylococcus epidermidis antimicrobial delta-toxin (phenol-soluble modulin-gamma) cooperates with host antimicrobial peptides to kill group A Streptococcus. PLoS One. 2010 Jan; 5(1):e8557.
50. Laborel-Preneron E, Bianchi P, Boralevi F, Lehours P, Fraysse F, Morice-Picard F, et al. Effects of the Staphylococcus aureus and Staphylococcus epidermidis Secretomes Isolated from the Skin Microbiota of Atopic Children on CD4+ T Cell Activation. PLoS One. 2015 Oct;10(10):e0141067.
51. Lai Y, Di Nardo A, Nakatsuji T, Leichtle A, Yang Y, Cogen AL, et al. Commensal bacteria regulate Toll-like receptor 3-dependent in- flammation after skin injury. Nat Med. 2009 Dec;15(12):1377–82.
52. Naik S, Bouladoux N, Wilhelm C, Molloy MJ, Salcedo R, Kastenmuller W, et al. Compart-mentalized control of skin immunity by resident commensals. Science. 2012 Aug; 337(6098):1115–9.
53. Naik S, Bouladoux N, Linehan JL, Han SJ, Harrison OJ, Wilhelm C, et al. Commensal- dendritic-cell interaction specifies a unique protective skin immune signature. Nature. 2015 Apr;520(7545):104–8.
54. Nakatsuji T, Chen TH, Butcher AM, Trzoss LL, Nam SJ, Shirakawa KT, et al. A commensal strain of Staphylococcus epidermidis protects against skin neoplasia. Sci Adv. 2018 Feb;4(2):eaao4502.
55. Wang Y, Zhang L, Yu J, Huang S, Wang Z, Chun KA, et al. A Co-Drug of Butyric Acid Derived from Fermentation Metabolites of the Human Skin Microbiome Stimulates Ad- ipogenic Differentiation of Adipose-Derived Stem Cells: Implications in Tissue Augmentation. J Invest Dermatol. 2017 Jan;137(1):46– 56.
56. Wang Y, Kuo S, Shu M, Yu J, Huang S, Dai A, et al. Staphylococcus epidermidis in the human skin microbiome mediates fermentation to inhibit the growth of Propionibacterium acnes: implications of probiotics in acne vulgaris. Appl Microbiol Biotechnol. 2014 Jan; 98(1):411–24.
57. Nishijima S, Kurokawa I, Katoh N, Watanabe K. The bacteriology of acne vulgaris and anti-microbial susceptibility of Propionibacterium acnes and Staphylococcus epidermidis isolated from acne lesions. J Dermatol. 2000 May; 27(5):318–23.
58. Skabytska Y, Biedermann T. Staphylococcus epidermidis Sets Things Right Again. J Invest Dermatol. 2016 Mar;136(3):559–60.
59. Xia X, Li Z, Liu K, Wu Y, Jiang D, Lai Y. Staph- ylococcal LTA-Induced miR-143 Inhibits Propionibacterium acnes Mediated Inflammatory Response in Skin. J Invest Dermatol. 2016 Mar;136(3):621–30.
60. Lai Y, Cogen AL, Radek KA, Park HJ, Macleod DT, Leichtle A, et al. Activation of TLR2 by a small molecule produced by Staphylococcus epidermidis increases antimicrobial– defense against bacterial skin infections. J Invest Dermatol. 2010 Sep;130(9):2211–21.
61. Bialecka A, Mak M, Biedron R, Bobek M, Kasprowicz A, Marcinkiewicz J. Different pro-inflammatory and immunogenic potentials of Propionibacterium acnes and Staphylococcus epidermidis: implications for chronic inflammatory acne. Arch Immunol Ther Exp (Warsz). 2005 Jan-Feb;53(1):79–85.
62. Michel S, Jomard A, Demarchez M. Pharmacology of adapalene. Br J Dermatol. 1998 Oct;139 Suppl 52:3–7.
63. Tenaud I, Khammari A, Dreno B. In vitro modulation of TLR-2, CD1d and IL-10 by adapalene on normal human skin and acne inflammatory lesions. Exp Dermatol. 2007 Jun;16(6):500–6.
64. Ozolins M, Eady EA, Avery AJ, Cunliffe WJ, Po AL, O’Neill C, et al. Comparison of five antimicrobial regimens for treatment of mild to moderate inflammatory facial acne vulgaris in the community: randomised controlled trial. Lancet. 2004 Dec;364(9452):2188–95.
65. Fulton JE Jr, Farzad-Bakshandeh A, Bradley S. Studies on the mechanism of action to topical benzoyl peroxide and vitamin A acid in acne vulgaris. J Cutan Pathol. 1974;1(5):191–200.
66. Moon SH, Roh HS, Kim YH, Kim JE, Ko JY, Ro YS. Antibiotic resistance of microbial strains isolated from Korean acne patients. J Dermatol. 2012 Oct;39(10):833–7.
67. Simonart T, Dramaix M, De Maertelaer V. Efficacy of tetracyclines in the treatment of acne vulgaris: a review. Br J Dermatol. 2008 Feb;158(2):208–16.
68. Leyden JJ, Del Rosso JQ, Baum EW. The use of isotretinoin in the treatment of acne vulgaris: clinical considerations and future directions. J Clin Aesthet Dermatol. 2014 Feb;7(2 Suppl):S3–21.
69. Ozlu E, Karadag AS, Ozkanli S, Oguztuzun S, Kilic M, Zemheri E, et al. Comparison of TLR-2, TLR-4, and antimicrobial peptide levels in different lesions of acne vulgaris. Cutan Ocul Toxicol. 2016 Dec;35(4):300–9.
70. Marcinkiewicz M, Majewski S. The role of antimicrobial peptides in chronic inflammatory skin diseases. Postepy Dermatol Alergol. 2016 Feb;33(1):6–12.
71. Feldman SR, Tan J, Poulin Y, Dirschka T, Kerrouche N, Manna V. The efficacy of adapalene-benzoyl peroxide combination increases with number of acne lesions. J Am Acad Dermatol. 2011 Jun;64(6):1085–91.
72. Jugeau S, Tenaud I, Knol AC, Jarrousse V, Quereux G, Khammari A, et al. Induction of toll-like receptors by Propionibacterium acnes. Br J Dermatol. 2005 Dec;153(6):1105–13.
73. Zaenglein AL, Pathy AL, Schlosser BJ, Alikhan A, Baldwin HE, Berson DS, et al. Guidelines of care for the management of acne vulgaris. J Am Acad Dermatol. 2016 May;74(5):945–73.e33.
74. Bowe W, Patel NB, Logan AC. Acne vulgaris, probiotics and the gut-brain-skin axis: from anecdote to translational medicine. Benef Microbes. 2014 Jun;5(2):185–99.
75. Arck P, Handjiski B, Hagen E, Pincus M, Bruenahl C, Bienenstock J, et al. Is there a ‘gut-brain-skin axis’? Exp Dermatol. 2010 May;19(5):401–5.
76. Gueniche A, Philippe D, Bastien P, Reuteler G, Blum S, Castiel-Higounenc I, et al. Randomised double-blind placebo-controlled study of the effect of Lactobacillus paracasei NCC 2461 on skin reactivity. Benef Microbes. 2014 Jun:5(2):137-45.
77. Gueniche A, Benyacoub J, Philippe D, Bastien P, Kusy N, Breton L, et al. Lactobacillus paracasei CNCM I-2116 (ST11) inhibits substance P-induced skin inflammation and accelerates skin barrier function recovery in vitro. Eur J Dermatol. 2010 Nov-Dec;20(6):731–7.
78. Fabbrocini G, Bertona M, Picazo O, Pareja-Galeano H, Monfrecola G, Emanuele E. Supplementation with Lactobacillus rhamnosus SP1 normalises skin expression of genes implicated in insulin signalling and improves adult acne. Benef Microbes. 2016 Nov;7(5):625–30.
79. Fuchs-Tarlovsky V, Marquez-Barba MF, Sriram K. Probiotics in dermatologic practice. Nutrition. 2016 Mar;32(3):289–95.
80. Baquerizo Nole KL, Yim E, Keri JE. Probiotics and prebiotics in dermatology. J Am Acad Dermatol. 2014 Oct;71(4):814–21.
81. Huang R, Ning H, Shen M, Li J, Zhang J, Chen X. Probiotics for the Treatment of Atopic Dermatitis in Children: A Systematic Review and Meta-Analysis of Randomized Controlled Trials. Front Cell Infect Microbiol. 2017 Sep;7:392.
82. Thio HB. The Microbiome in Psoriasis and Psoriatic Arthritis: The Skin Perspective. J Rheumatol Suppl. 2018 Jun;94:30–1.
83. Chen YH, Wu CS, Chao YH, Lin CC, Tsai HY, Li YR, et al. Lactobacillus pentosus GMNL-77 inhibits skin lesions in imiquimod-induced psoriasis-like mice. Yao Wu Shi Pin Fen Xi. 2017 Jul;25(3):559–66.
84. Blank-Porat D, Gruss-Fischer T, Tarasenko N, Malik Z, Nudelman A, Rephaeli A. The anticancer prodrugs of butyric acid AN-7 and AN-9, possess antiangiogenic properties. Cancer Lett. 2007 Oct;256(1):39–48.
Mots-clés: Dermatologie clinique & chirurgie dermatologique
Partagez cet article sur




